Workflow Type: Galaxy
Open
Frozen
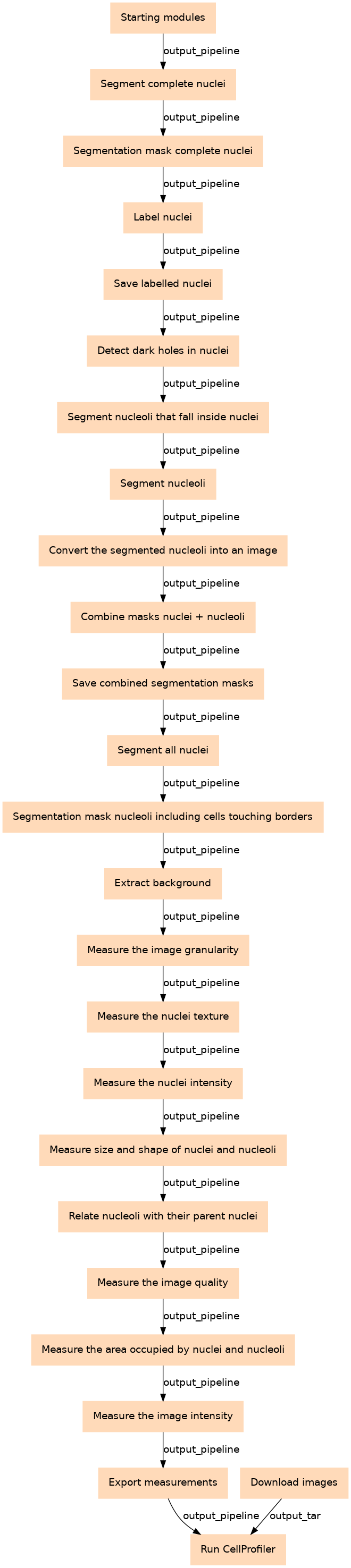
Nucleoli segmentation and feature extraction using CellProfiler
Associated Tutorial
This workflows is part of the tutorial CP_pipeline_IDR_training, available in the GTN
Thanks to...
Tutorial Author(s): Beatriz Serrano-Solano, Jean-Karim Hériché
Steps
| ID | Name | Description |
|---|---|---|
| 0 | Starting modules | Detail the metadata associated to the images that will be processed toolshed.g2.bx.psu.edu/repos/bgruening/cp_common/cp_common/3.1.9 |
| 1 | Download images | toolshed.g2.bx.psu.edu/repos/iuc/idr_download_by_ids/idr_download_by_ids/0.41 |
| 2 | Segment complete nuclei | Incomplete nuclei that are touching the borders of the image are ignored and therefore, not segmented. It doesn't include the nuclei outside the diameter range either. toolshed.g2.bx.psu.edu/repos/bgruening/cp_identify_primary_objects/cp_identify_primary_objects/3.1.9 |
| 3 | Segmentation mask complete nuclei | toolshed.g2.bx.psu.edu/repos/bgruening/cp_convert_objects_to_image/cp_convert_objects_to_image/3.1.9 |
| 4 | Label nuclei | toolshed.g2.bx.psu.edu/repos/bgruening/cp_display_data_on_image/cp_display_data_on_image/3.1.9 |
| 5 | Save labelled nuclei | toolshed.g2.bx.psu.edu/repos/bgruening/cp_save_images/cp_save_images/3.1.9 |
| 6 | Detect dark holes in nuclei | toolshed.g2.bx.psu.edu/repos/bgruening/cp_enhance_or_suppress_features/cp_enhance_or_suppress_features/3.1.9 |
| 7 | Segment nucleoli that fall inside nuclei | The dark holes enhanced in the previous step are segmented only if they are inside a nuclei toolshed.g2.bx.psu.edu/repos/bgruening/cp_mask_image/cp_mask_image/3.1.9 |
| 8 | Segment nucleoli | The segmentation only affects those inside the nuclei toolshed.g2.bx.psu.edu/repos/bgruening/cp_identify_primary_objects/cp_identify_primary_objects/3.1.9 |
| 9 | Convert the segmented nucleoli into an image | toolshed.g2.bx.psu.edu/repos/bgruening/cp_convert_objects_to_image/cp_convert_objects_to_image/3.1.9 |
| 10 | Combine masks (nuclei + nucleoli) | toolshed.g2.bx.psu.edu/repos/bgruening/cp_gray_to_color/cp_gray_to_color/3.1.9 |
| 11 | Save combined segmentation masks | toolshed.g2.bx.psu.edu/repos/bgruening/cp_save_images/cp_save_images/3.1.9 |
| 12 | Segment all nuclei | Incomplete nuclei that are touching the borders of the image are segmented, also the nuclei outside the diameter range. toolshed.g2.bx.psu.edu/repos/bgruening/cp_identify_primary_objects/cp_identify_primary_objects/3.1.9 |
| 13 | Segmentation mask nucleoli including cells touching borders | toolshed.g2.bx.psu.edu/repos/bgruening/cp_convert_objects_to_image/cp_convert_objects_to_image/3.1.9 |
| 14 | Extract background | toolshed.g2.bx.psu.edu/repos/bgruening/cp_image_math/cp_image_math/0.1.0 |
| 15 | Measure the image granularity | This step measures the granularity of the original image toolshed.g2.bx.psu.edu/repos/bgruening/cp_measure_granularity/cp_measure_granularity/3.1.9 |
| 16 | Measure the nuclei texture | This step measures the texture of the original image and nuclei in it toolshed.g2.bx.psu.edu/repos/bgruening/cp_measure_texture/cp_measure_texture/3.1.9 |
| 17 | Measure the nuclei intensity | This step measures the intensity of the original image and the nuclei toolshed.g2.bx.psu.edu/repos/bgruening/cp_measure_object_intensity/cp_measure_object_intensity/3.1.9 |
| 18 | Measure size and shape of nuclei and nucleoli | toolshed.g2.bx.psu.edu/repos/bgruening/cp_measure_object_size_shape/cp_measure_object_size_shape/3.1.9 |
| 19 | Relate nucleoli with their parent nuclei | toolshed.g2.bx.psu.edu/repos/bgruening/cp_relate_objects/cp_relate_objects/3.1.9 |
| 20 | Measure the image quality | toolshed.g2.bx.psu.edu/repos/bgruening/cp_measure_image_quality/cp_measure_image_quality/3.1.9 |
| 21 | Measure the area occupied by nuclei and nucleoli | toolshed.g2.bx.psu.edu/repos/bgruening/cp_measure_image_area_occupied/cp_measure_image_area_occupied/3.1.9 |
| 22 | Measure the image intensity | toolshed.g2.bx.psu.edu/repos/bgruening/cp_measure_image_intensity/cp_measure_image_intensity/3.1.9 |
| 23 | Export measurements | toolshed.g2.bx.psu.edu/repos/bgruening/cp_export_to_spreadsheet/cp_export_to_spreadsheet/3.1.9 |
| 24 | Run CellProfiler | toolshed.g2.bx.psu.edu/repos/bgruening/cp_cellprofiler/cp_cellprofiler/3.1.9 |
Outputs
| ID | Name | Description | Type |
|---|---|---|---|
| _anonymous_output_1 | #main/_anonymous_output_1 | n/a |
|
| _anonymous_output_10 | #main/_anonymous_output_10 | n/a |
|
| _anonymous_output_11 | #main/_anonymous_output_11 | n/a |
|
| _anonymous_output_12 | #main/_anonymous_output_12 | n/a |
|
| _anonymous_output_13 | #main/_anonymous_output_13 | n/a |
|
| _anonymous_output_14 | #main/_anonymous_output_14 | n/a |
|
| _anonymous_output_15 | #main/_anonymous_output_15 | n/a |
|
| _anonymous_output_16 | #main/_anonymous_output_16 | n/a |
|
| _anonymous_output_17 | #main/_anonymous_output_17 | n/a |
|
| _anonymous_output_18 | #main/_anonymous_output_18 | n/a |
|
| _anonymous_output_19 | #main/_anonymous_output_19 | n/a |
|
| _anonymous_output_2 | #main/_anonymous_output_2 | n/a |
|
| _anonymous_output_20 | #main/_anonymous_output_20 | n/a |
|
| _anonymous_output_21 | #main/_anonymous_output_21 | n/a |
|
| _anonymous_output_22 | #main/_anonymous_output_22 | n/a |
|
| _anonymous_output_23 | #main/_anonymous_output_23 | n/a |
|
| _anonymous_output_24 | #main/_anonymous_output_24 | n/a |
|
| _anonymous_output_25 | #main/_anonymous_output_25 | n/a |
|
| _anonymous_output_26 | #main/_anonymous_output_26 | n/a |
|
| _anonymous_output_3 | #main/_anonymous_output_3 | n/a |
|
| _anonymous_output_4 | #main/_anonymous_output_4 | n/a |
|
| _anonymous_output_5 | #main/_anonymous_output_5 | n/a |
|
| _anonymous_output_6 | #main/_anonymous_output_6 | n/a |
|
| _anonymous_output_7 | #main/_anonymous_output_7 | n/a |
|
| _anonymous_output_8 | #main/_anonymous_output_8 | n/a |
|
| _anonymous_output_9 | #main/_anonymous_output_9 | n/a |
|
Version History
3.0 (latest) Created 16th Jul 2024 at 14:10 by Helena Rasche
Added/updated 4 files
Open
 master
master88cf200
7.0 (earliest) Created 25th Jun 2024 at 11:16 by Helena Rasche
Added/updated 4 files
Frozen
 7.0
7.04c11662
 Creators and Submitter
Creators and SubmitterCreators
Not specifiedSubmitter
Discussion Channel
Tool
Activity
Views: 615 Downloads: 181
Created: 25th Jun 2024 at 11:16
Last updated: 25th Jun 2024 at 11:16
 Tags
Tags Attributions
AttributionsNone
 Visit source
Visit source
 https://orcid.org/0000-0001-9760-8992
https://orcid.org/0000-0001-9760-8992