Workflow Type: Galaxy
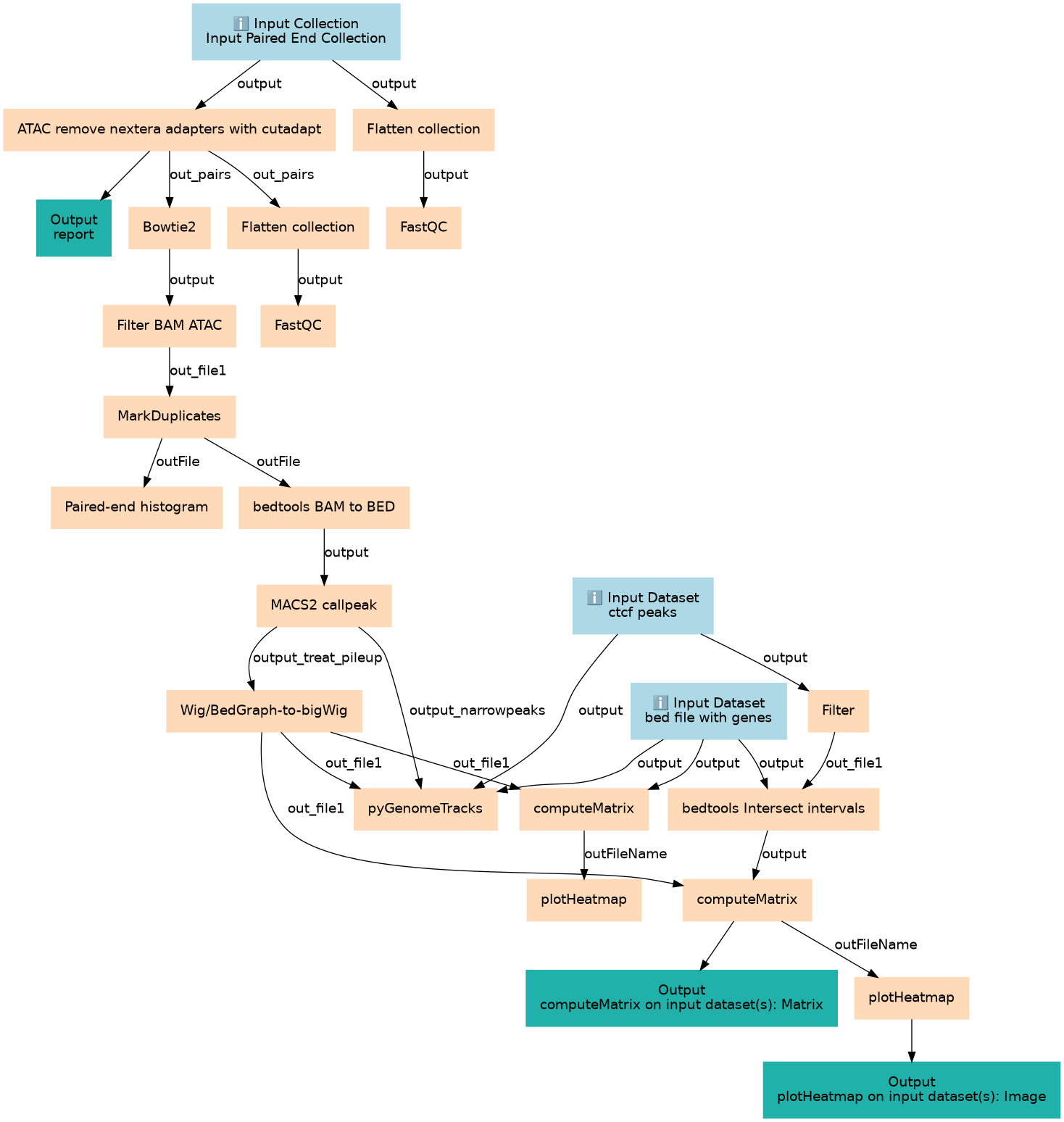
atac-seq
Associated Tutorial
This workflows is part of the tutorial ATAC-Seq data analysis, available in the GTN
Features
- Includes Galaxy Workflow Tests
- Includes a Galaxy Workflow Report
Thanks to...
Workflow Author(s): Tristan Reynolds
Tutorial Author(s): Lucille Delisle, Maria Doyle, Florian Heyl
Tutorial Contributor(s): Tristan Reynolds, Helena Rasche, Saskia Hiltemann, Yvan Le Bras, Björn Grüning, Nicola Soranzo, Lucille Delisle, Gildas Le Corguillé, Hans-Rudolf Hotz
Funder(s): The University of Melbourne, Melbourne Bioinformatics, Australian BioCommons
Inputs
| ID | Name | Description | Type |
|---|---|---|---|
| Input Paired End Collection | Input Paired End Collection | n/a |
|
| bed file with genes | bed file with genes | n/a |
|
| ctcf peaks | ctcf peaks | n/a |
|
Steps
| ID | Name | Description |
|---|---|---|
| 3 | ATAC remove nextera adapters with cutadapt | toolshed.g2.bx.psu.edu/repos/lparsons/cutadapt/cutadapt/5.1+galaxy0 |
| 4 | Flatten collection | __FLATTEN__ |
| 5 | Filter | Filter1 |
| 6 | Bowtie2 | toolshed.g2.bx.psu.edu/repos/devteam/bowtie2/bowtie2/2.4.2+galaxy0 |
| 7 | Flatten collection | __FLATTEN__ |
| 8 | FastQC | toolshed.g2.bx.psu.edu/repos/devteam/fastqc/fastqc/0.72+galaxy1 |
| 9 | bedtools Intersect intervals | toolshed.g2.bx.psu.edu/repos/iuc/bedtools/bedtools_intersectbed/2.30.0 |
| 10 | Filter BAM ATAC | toolshed.g2.bx.psu.edu/repos/devteam/bamtools_filter/bamFilter/2.4.1 |
| 11 | FastQC | toolshed.g2.bx.psu.edu/repos/devteam/fastqc/fastqc/0.72+galaxy1 |
| 12 | MarkDuplicates | toolshed.g2.bx.psu.edu/repos/devteam/picard/picard_MarkDuplicates/2.18.2.2 |
| 13 | Paired-end histogram | toolshed.g2.bx.psu.edu/repos/iuc/pe_histogram/pe_histogram/1.0.1 |
| 14 | bedtools BAM to BED | toolshed.g2.bx.psu.edu/repos/iuc/bedtools/bedtools_bamtobed/2.30.0 |
| 15 | MACS2 callpeak | toolshed.g2.bx.psu.edu/repos/iuc/macs2/macs2_callpeak/2.1.1.20160309.6 |
| 16 | Wig/BedGraph-to-bigWig | wig_to_bigWig |
| 17 | pyGenomeTracks | toolshed.g2.bx.psu.edu/repos/iuc/pygenometracks/pygenomeTracks/3.6 |
| 18 | computeMatrix | toolshed.g2.bx.psu.edu/repos/bgruening/deeptools_compute_matrix/deeptools_compute_matrix/3.3.2.0.0 |
| 19 | computeMatrix | toolshed.g2.bx.psu.edu/repos/bgruening/deeptools_compute_matrix/deeptools_compute_matrix/3.3.2.0.0 |
| 20 | plotHeatmap | toolshed.g2.bx.psu.edu/repos/bgruening/deeptools_plot_heatmap/deeptools_plot_heatmap/3.3.2.0.1 |
| 21 | plotHeatmap | toolshed.g2.bx.psu.edu/repos/bgruening/deeptools_plot_heatmap/deeptools_plot_heatmap/3.3.2.0.1 |
Outputs
| ID | Name | Description | Type |
|---|---|---|---|
| report | report | n/a |
|
| _anonymous_output_1 | _anonymous_output_1 | n/a |
|
| _anonymous_output_2 | _anonymous_output_2 | n/a |
|
| _anonymous_output_3 | _anonymous_output_3 | n/a |
|
| _anonymous_output_4 | _anonymous_output_4 | n/a |
|
| _anonymous_output_5 | _anonymous_output_5 | n/a |
|
| _anonymous_output_6 | _anonymous_output_6 | n/a |
|
| _anonymous_output_7 | _anonymous_output_7 | n/a |
|
| _anonymous_output_8 | _anonymous_output_8 | n/a |
|
| _anonymous_output_9 | _anonymous_output_9 | n/a |
|
| _anonymous_output_10 | _anonymous_output_10 | n/a |
|
| _anonymous_output_11 | _anonymous_output_11 | n/a |
|
| _anonymous_output_12 | _anonymous_output_12 | n/a |
|
| _anonymous_output_13 | _anonymous_output_13 | n/a |
|
| _anonymous_output_14 | _anonymous_output_14 | n/a |
|
| _anonymous_output_15 | _anonymous_output_15 | n/a |
|
| _anonymous_output_16 | _anonymous_output_16 | n/a |
|
| _anonymous_output_17 | _anonymous_output_17 | n/a |
|
| _anonymous_output_18 | _anonymous_output_18 | n/a |
|
| computeMatrix on input dataset(s): Matrix | computeMatrix on input dataset(s): Matrix | n/a |
|
| _anonymous_output_19 | _anonymous_output_19 | n/a |
|
| plotHeatmap on input dataset(s): Image | plotHeatmap on input dataset(s): Image | n/a |
|
Version History
 Creators and Submitter
Creators and SubmitterCreators
Not specifiedSubmitter
Discussion Channel
Activity
Views: 2295 Downloads: 358 Runs: 2
Created: 2nd Jun 2025 at 10:59
Last updated: 22nd Dec 2025 at 13:14
 Tags
Tags Attributions
AttributionsNone
 Visit source
Visit source Run on Galaxy
Run on Galaxy
 master
master 10.0
10.0