Workflow Type: Galaxy
Associated Tutorial
This workflows is part of the tutorial Remove contamination and host reads, available in the GTN
Features
- Includes Galaxy Workflow Tests
- Includes a Galaxy Workflow Report
Thanks to...
Workflow Author(s): Paul Zierep
Tutorial Author(s): Mina Hojat Ansari, Bérénice Batut
Inputs
| ID | Name | Description | Type |
|---|---|---|---|
| Input paired fastq | Input paired fastq | n/a |
|
| Reference Genome Build In | Reference Genome Build In | n/a |
|
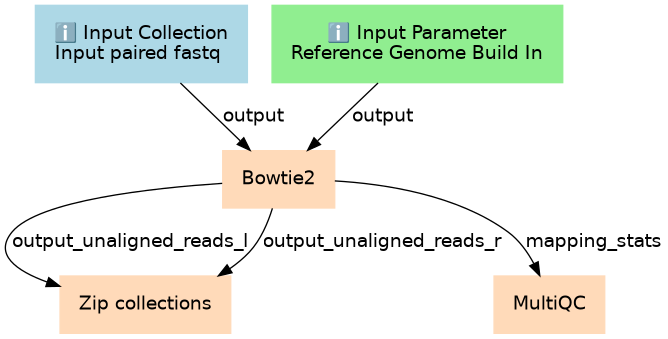
Steps
| ID | Name | Description |
|---|---|---|
| 2 | Bowtie2 | toolshed.g2.bx.psu.edu/repos/devteam/bowtie2/bowtie2/2.5.3+galaxy1 |
| 3 | Zip collections | __ZIP_COLLECTION__ |
| 4 | MultiQC | toolshed.g2.bx.psu.edu/repos/iuc/multiqc/multiqc/1.27+galaxy3 |
Version History
 Creators and Submitter
Creators and SubmitterCreators
Not specifiedSubmitter
Discussion Channel
License
Activity
Views: 758 Downloads: 106 Runs: 0
Created: 22nd Dec 2025 at 13:13
 Tags
Tags Attributions
AttributionsNone
 Visit source
Visit source Run on Galaxy
Run on Galaxy
 master
master