Workflow Type: Galaxy
Associated Tutorial
This workflows is part of the tutorial Segmentation of Anatomical Structures in Medical 3-D Images, available in the GTN
Features
- Includes Galaxy Workflow Tests
- Includes a Galaxy Workflow Report
- Uses Galaxy Workflow Comments
Thanks to...
Workflow Author(s): Leonid Kostrykin
Tutorial Author(s): Leonid Kostrykin
Tutorial Contributor(s): Diana Chiang Jurado, Beatriz Serrano-Solano, Leonid Kostrykin, Saskia Hiltemann
Funder(s): German Network for Bioinformatics Infrastructure Service, Training, Cooperations & Cloud Computing
Inputs
| ID | Name | Description | Type |
|---|---|---|---|
| bones_threshold_huv | bones_threshold_huv | Bones Threshold Value (HU) |
|
| dicom_series | dicom_series | DICOM Series |
|
| y_tol_bones_px | y_tol_bones_px | Y-Tolerance for Bones Segmentation (pixel) |
|
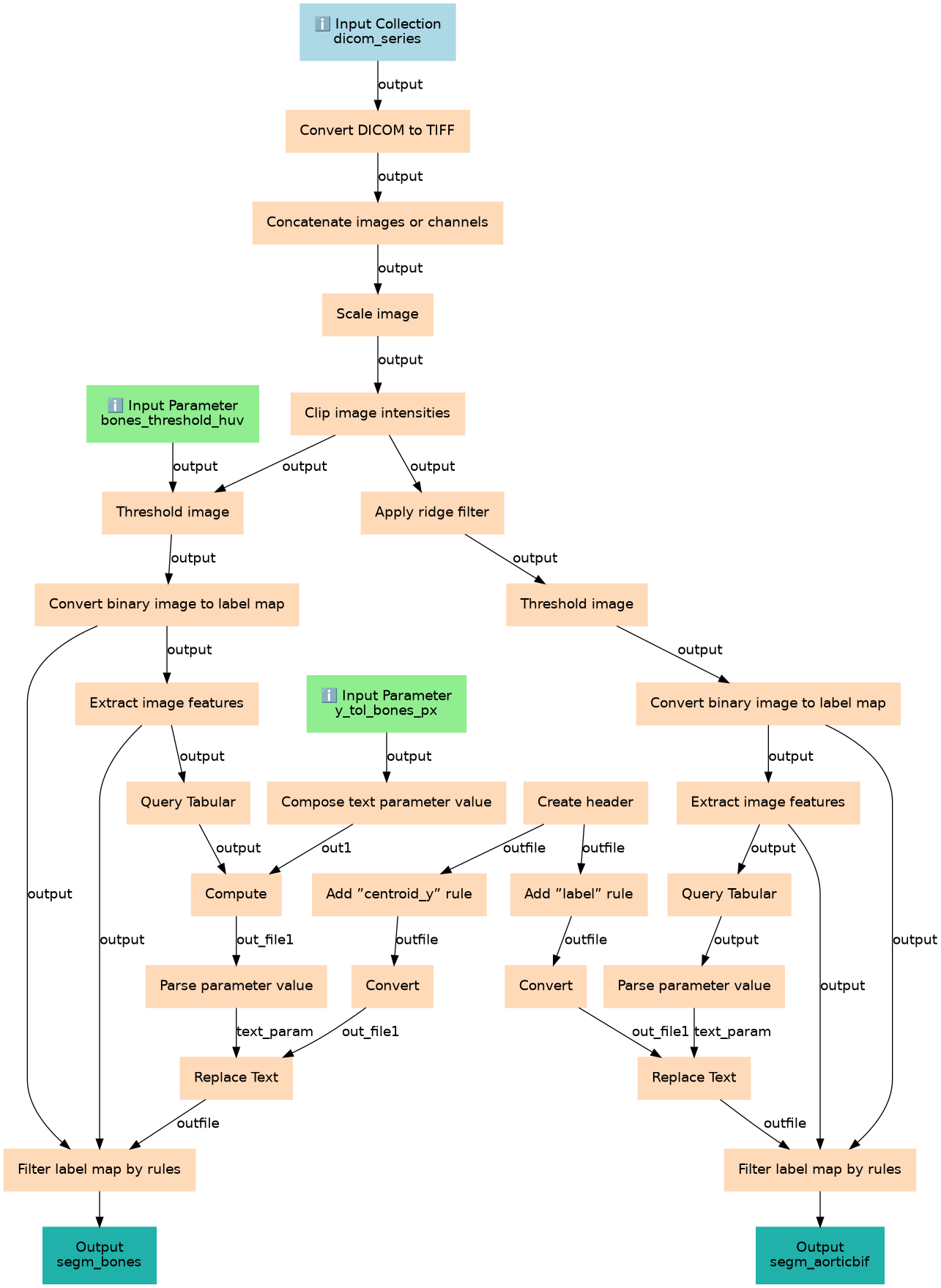
Steps
| ID | Name | Description |
|---|---|---|
| 3 | Create header | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_text_file_with_recurring_lines/9.5+galaxy2 |
| 4 | Compose text parameter value | toolshed.g2.bx.psu.edu/repos/iuc/compose_text_param/compose_text_param/0.1.1 |
| 5 | Convert DICOM to TIFF | toolshed.g2.bx.psu.edu/repos/imgteam/highdicom_dicom2tiff/highdicom_dicom2tiff/0.27.0+galaxy0 |
| 6 | Add "centroid_y" rule | Add row toolshed.g2.bx.psu.edu/repos/bgruening/add_line_to_file/add_line_to_file/0.1.0 |
| 7 | Add "label" rule | toolshed.g2.bx.psu.edu/repos/bgruening/add_line_to_file/add_line_to_file/0.1.0 |
| 8 | Concatenate images or channels | toolshed.g2.bx.psu.edu/repos/imgteam/concat_channels/ip_concat_channels/0.5 |
| 9 | Convert | Convert characters1 |
| 10 | Convert | Convert characters1 |
| 11 | Scale image | toolshed.g2.bx.psu.edu/repos/imgteam/scale_image/ip_scale_image/0.25.2+galaxy0 |
| 12 | Clip image intensities | toolshed.g2.bx.psu.edu/repos/imgteam/clip_image/clip_image/0.7.3+galaxy0 |
| 13 | Threshold image | toolshed.g2.bx.psu.edu/repos/imgteam/2d_auto_threshold/ip_threshold/0.25.2+galaxy0 |
| 14 | Apply ridge filter | toolshed.g2.bx.psu.edu/repos/imgteam/ridge_filter/ridge_filter_skimage/0.22.0+galaxy2 |
| 15 | Convert binary image to label map | toolshed.g2.bx.psu.edu/repos/imgteam/binary2labelimage/ip_binary_to_labelimage/0.7.3+galaxy0 |
| 16 | Threshold image | toolshed.g2.bx.psu.edu/repos/imgteam/2d_auto_threshold/ip_threshold/0.25.2+galaxy0 |
| 17 | Extract image features | toolshed.g2.bx.psu.edu/repos/imgteam/2d_feature_extraction/ip_2d_feature_extraction/0.25.2+galaxy1 |
| 18 | Convert binary image to label map | toolshed.g2.bx.psu.edu/repos/imgteam/binary2labelimage/ip_binary_to_labelimage/0.7.3+galaxy0 |
| 19 | Query Tabular | toolshed.g2.bx.psu.edu/repos/iuc/query_tabular/query_tabular/3.3.2 |
| 20 | Extract image features | toolshed.g2.bx.psu.edu/repos/imgteam/2d_feature_extraction/ip_2d_feature_extraction/0.25.2+galaxy1 |
| 21 | Compute | toolshed.g2.bx.psu.edu/repos/devteam/column_maker/Add_a_column1/2.1 |
| 22 | Query Tabular | toolshed.g2.bx.psu.edu/repos/iuc/query_tabular/query_tabular/3.3.2 |
| 23 | Parse parameter value | param_value_from_file |
| 24 | Parse parameter value | param_value_from_file |
| 25 | Replace Text | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_replace_in_line/9.5+galaxy2 |
| 26 | Replace Text | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_replace_in_line/9.5+galaxy2 |
| 27 | Filter label map by rules | toolshed.g2.bx.psu.edu/repos/imgteam/2d_filter_segmentation_by_features/ip_2d_filter_segmentation_by_features/0.7.3+galaxy1 |
| 28 | Filter label map by rules | toolshed.g2.bx.psu.edu/repos/imgteam/2d_filter_segmentation_by_features/ip_2d_filter_segmentation_by_features/0.7.3+galaxy1 |
Outputs
| ID | Name | Description | Type |
|---|---|---|---|
| segm_bones | segm_bones | n/a |
|
| segm_aorticbif | segm_aorticbif | n/a |
|
Version History
 Creators and Submitter
Creators and SubmitterCreators
Not specifiedSubmitter
Discussion Channel
License
Activity
Views: 270 Downloads: 59 Runs: 0
Created: 9th Mar 2026 at 13:26
 Attributions
AttributionsNone
 Visit source
Visit source Run on Galaxy
Run on Galaxy
 master
master