Workflow Type: Galaxy
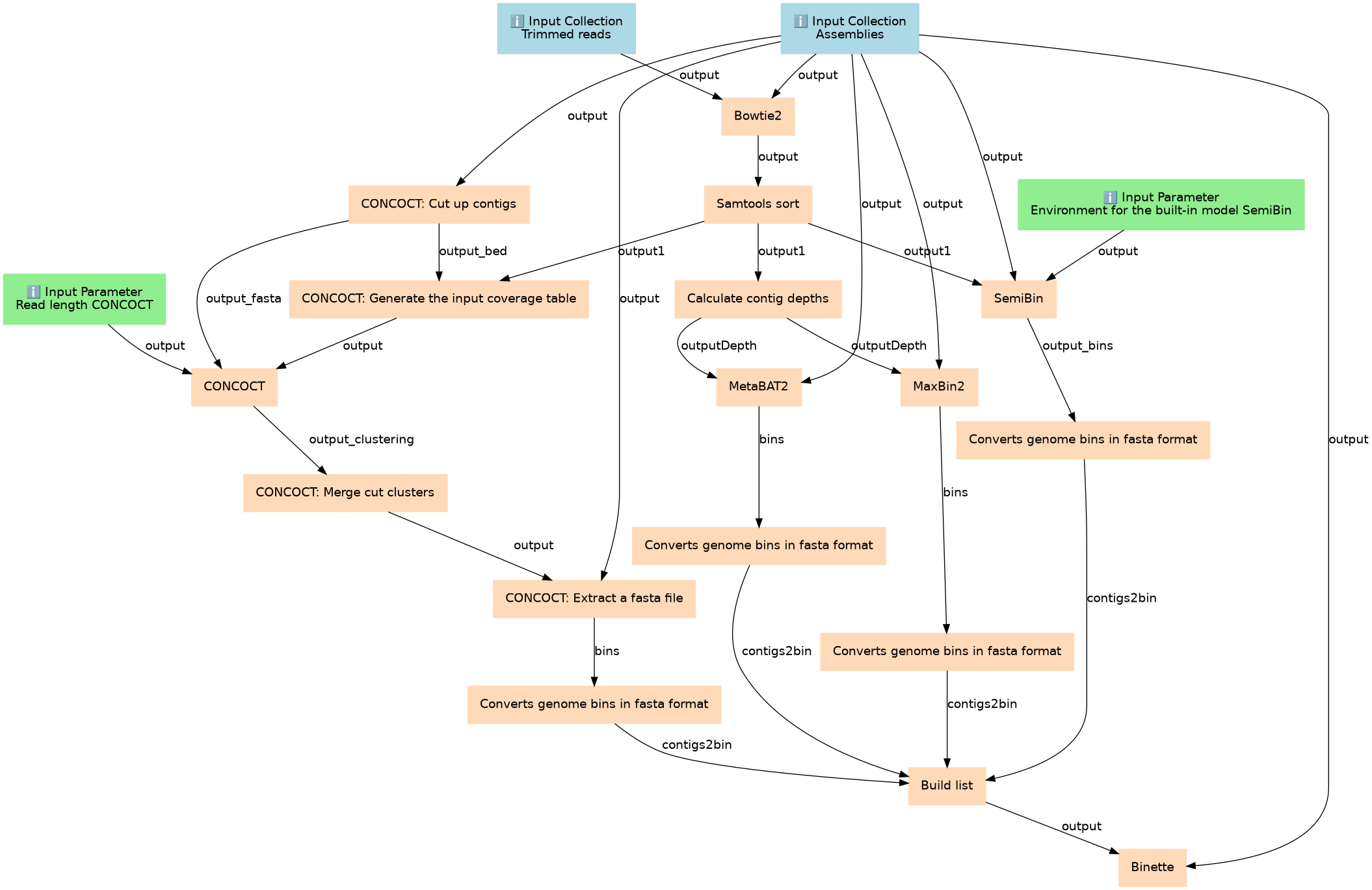
Binning workflows that uses abundance information and performs binning of metagenomic contigs using 4 different binners as well as bin refinement.
Associated Tutorial
This workflows is part of the tutorial Binning of metagenomic sequencing data, available in the GTN
Features
- Includes Galaxy Workflow Tests
- Includes a Galaxy Workflow Report
- Uses Galaxy Workflow Comments
Thanks to...
Workflow Author(s): Paul Zierep
Tutorial Author(s): Paul Zierep, Nikos Pechlivanis, Fotis E. Psomopoulos, Vini Salazar
Tutorial Contributor(s): Bérénice Batut, Teresa Müller, Deepti Varshney, Nikos Pechlivanis, Helena Rasche, Saskia Hiltemann
Inputs
| ID | Name | Description | Type |
|---|---|---|---|
| Assemblies | Assemblies | This workflow allows using a custom assembly as input. If provided, select `custom assembly` as Assembler. Provide one assembly for each group of trimmed input reads. |
|
| Environment for the built-in model (SemiBin) | Environment for the built-in model (SemiBin) | Environment for the built-in model (SemiBin), options are: human_gut, dog_gut, ocean, soil, cat_gut, human_oral, mouse_gut, pig_gut, built_environment, wastewater, chicken_caecum, global |
|
| Read length (CONCOCT) | Read length (CONCOCT) | CONCOCT requires the read length for coverage. Best use fastQC to estimate the mean value. |
|
| Trimmed reads | Trimmed reads | Samples grouped for co-assembly. For individual assembly use same reads as `Trimmed reads input`. The tool fastq_groupmerge can be used to perform the grouping. |
|
Steps
| ID | Name | Description |
|---|---|---|
| 4 | CONCOCT: Cut up contigs | toolshed.g2.bx.psu.edu/repos/iuc/concoct_cut_up_fasta/concoct_cut_up_fasta/1.1.0+galaxy2 |
| 5 | Bowtie2 | toolshed.g2.bx.psu.edu/repos/devteam/bowtie2/bowtie2/2.5.4+galaxy0 |
| 6 | Samtools sort | toolshed.g2.bx.psu.edu/repos/devteam/samtools_sort/samtools_sort/2.0.7 |
| 7 | CONCOCT: Generate the input coverage table | toolshed.g2.bx.psu.edu/repos/iuc/concoct_coverage_table/concoct_coverage_table/1.1.0+galaxy2 |
| 8 | Calculate contig depths | toolshed.g2.bx.psu.edu/repos/iuc/metabat2_jgi_summarize_bam_contig_depths/metabat2_jgi_summarize_bam_contig_depths/2.17+galaxy0 |
| 9 | SemiBin | toolshed.g2.bx.psu.edu/repos/iuc/semibin/semibin/2.1.0+galaxy1 |
| 10 | CONCOCT | toolshed.g2.bx.psu.edu/repos/iuc/concoct/concoct/1.1.0+galaxy2 |
| 11 | MetaBAT2 | toolshed.g2.bx.psu.edu/repos/iuc/metabat2/metabat2/2.17+galaxy0 |
| 12 | MaxBin2 | toolshed.g2.bx.psu.edu/repos/mbernt/maxbin2/maxbin2/2.2.7+galaxy6 |
| 13 | Converts genome bins in fasta format | toolshed.g2.bx.psu.edu/repos/iuc/fasta_to_contig2bin/Fasta_to_Contig2Bin/1.1.7+galaxy1 |
| 14 | CONCOCT: Merge cut clusters | toolshed.g2.bx.psu.edu/repos/iuc/concoct_merge_cut_up_clustering/concoct_merge_cut_up_clustering/1.1.0+galaxy2 |
| 15 | Converts genome bins in fasta format | toolshed.g2.bx.psu.edu/repos/iuc/fasta_to_contig2bin/Fasta_to_Contig2Bin/1.1.7+galaxy1 |
| 16 | Converts genome bins in fasta format | toolshed.g2.bx.psu.edu/repos/iuc/fasta_to_contig2bin/Fasta_to_Contig2Bin/1.1.7+galaxy1 |
| 17 | CONCOCT: Extract a fasta file | toolshed.g2.bx.psu.edu/repos/iuc/concoct_extract_fasta_bins/concoct_extract_fasta_bins/1.1.0+galaxy2 |
| 18 | Converts genome bins in fasta format | toolshed.g2.bx.psu.edu/repos/iuc/fasta_to_contig2bin/Fasta_to_Contig2Bin/1.1.7+galaxy1 |
| 19 | Build list | __BUILD_LIST__ |
| 20 | Binette | toolshed.g2.bx.psu.edu/repos/iuc/binette/binette/1.2.0+galaxy0 |
Version History
 Creators and Submitter
Creators and SubmitterCreators
Not specifiedSubmitter
Discussion Channel
License
Activity
Views: 873 Downloads: 134 Runs: 0
Created: 8th Dec 2025 at 13:14
 Attributions
AttributionsNone
 Visit source
Visit source Run on Galaxy
Run on Galaxy
 master
master